diff --git a/.gitignore b/.gitignore
index d7d7bff..0c518ae 100644
--- a/.gitignore
+++ b/.gitignore
@@ -2,3 +2,8 @@
*.swp
__pycache__
tmp.py
+.env
+*.aux
+*.log
+*.out
+tmp.*
diff --git a/Makefile b/Makefile
new file mode 100644
index 0000000..58f2d1f
--- /dev/null
+++ b/Makefile
@@ -0,0 +1,11 @@
+.PHONY:to_wwu
+
+all: to_wwu
+
+to_wwu:
+ rsync -avz ~/577/repo/docs/figures/ linux-04:/home/jensen33/Dev/studentweb/assets/static/577/
+ scp ~/577/repo/docs/presentation.md linux-04:/home/jensen33/Dev/studentweb/content/577/contents.lr
+ scp ~/Dev/www.publicmatt.com/models/slides.ini linux-04:/home/jensen33/Dev/studentweb/models/
+ scp ~/Dev/www.publicmatt.com/templates/slides.html linux-04:/home/jensen33/Dev/studentweb/templates/
+ rsync -avz ~/Dev/www.publicmatt.com/assets/static/revealjs linux-04:/home/jensen33/Dev/studentweb/assets/static/
+ ssh linux-04 cd /home/jensen33/Dev/studentweb \; make
diff --git a/docs/Makefile b/docs/Makefile
new file mode 100644
index 0000000..a63e81e
--- /dev/null
+++ b/docs/Makefile
@@ -0,0 +1,3 @@
+paper.pdf: paper.tex
+ pdflatex $^ -o $@
+ evince $@
diff --git a/docs/figures/allsides_request.png b/docs/figures/allsides_request.png
new file mode 100644
index 0000000..920b104
Binary files /dev/null and b/docs/figures/allsides_request.png differ
diff --git a/docs/figures/articles_per_year.png b/docs/figures/articles_per_year.png
new file mode 100644
index 0000000..7a41821
Binary files /dev/null and b/docs/figures/articles_per_year.png differ
diff --git a/docs/figures/common_tld.png b/docs/figures/common_tld.png
new file mode 100644
index 0000000..e98b03f
Binary files /dev/null and b/docs/figures/common_tld.png differ
diff --git a/docs/figures/distinct_publishers.png b/docs/figures/distinct_publishers.png
new file mode 100644
index 0000000..e7113e2
Binary files /dev/null and b/docs/figures/distinct_publishers.png differ
diff --git a/docs/figures/link_cluster_elbow.png b/docs/figures/link_cluster_elbow.png
new file mode 100644
index 0000000..2764351
Binary files /dev/null and b/docs/figures/link_cluster_elbow.png differ
diff --git a/docs/figures/link_pca_clusters_links.png b/docs/figures/link_pca_clusters_links.png
new file mode 100644
index 0000000..edd7229
Binary files /dev/null and b/docs/figures/link_pca_clusters_links.png differ
diff --git a/docs/figures/link_pca_clusters_normalized.png b/docs/figures/link_pca_clusters_normalized.png
new file mode 100644
index 0000000..da0cc82
Binary files /dev/null and b/docs/figures/link_pca_clusters_normalized.png differ
diff --git a/docs/figures/link_pca_clusters_onehot.png b/docs/figures/link_pca_clusters_onehot.png
new file mode 100644
index 0000000..94941c0
Binary files /dev/null and b/docs/figures/link_pca_clusters_onehot.png differ
diff --git a/docs/figures/pca_with_classes.png b/docs/figures/pca_with_classes.png
new file mode 100644
index 0000000..a0362ef
Binary files /dev/null and b/docs/figures/pca_with_classes.png differ
diff --git a/docs/figures/stories_per_publisher.png b/docs/figures/stories_per_publisher.png
new file mode 100644
index 0000000..f63d983
Binary files /dev/null and b/docs/figures/stories_per_publisher.png differ
diff --git a/docs/figures/top_publishers.png b/docs/figures/top_publishers.png
new file mode 100644
index 0000000..8961cb7
Binary files /dev/null and b/docs/figures/top_publishers.png differ
diff --git a/docs/paper.pdf b/docs/paper.pdf
new file mode 100644
index 0000000..d374c95
Binary files /dev/null and b/docs/paper.pdf differ
diff --git a/docs/paper.tex b/docs/paper.tex
new file mode 100644
index 0000000..9e5af5d
--- /dev/null
+++ b/docs/paper.tex
@@ -0,0 +1,61 @@
+\documentclass{article}
+\usepackage{multicol}
+\usepackage{hyperref}
+\title{Data Mining CS 571}
+\author{Matt Jensen}
+\date{2023-04-25}
+
+\begin{document}
+\maketitle
+
+\section*{Abstract}
+
+News organizations have been repeatedly accused of being partisan.
+Additionally, they have been accused of polarizing dicussion to drive up revenue and engagement.
+This paper seeks to quantify those claims by classifying the degree to which news headlines have become more emotionally charged of time.
+A secondary goal is the investigate whether news organization have been uniformly polarized, or if one pole has been 'moving' more rapidly away from the 'middle'.
+This analysis will probe to what degree has the \href{https://en.wikipedia.org/wiki/Overton_window}{Overton Window} has shifted in the media.
+Naom Chomsky had a hypothesis about manufactured consent that is beyond the scope of this paper, so we will restrict our analysis to the presence of agenda instead of the cause of it.
+
+\begin{multicols}{2}
+
+\section{Data Preparation}
+The subject of analysis is a set of news article headlines scraped from the news aggregation site \href{https://mememorandum.com}{Memeorandum} for news stories from 2006 to 2022.
+Each news article has a title, author, description, publisher, publish date, url and related discussions.
+The site also has a concept of references, where a main, popular story may be covered by other sources.
+This link association might be used to support one or more of the hypothesis of the main analysis.
+After scraping the site, the data will need to be deduplicated and normalized to minimize storage costs and processing errors.
+What remains after these cleaning steps is approximitely 6,400 days of material, 300,000 distinct headlines from 21,000 publishers and 34,000 authors used in the study.
+
+\section{Missing Data Policy}
+
+The largest data policy that will have to be dealt with is news organizations that share the same parent company, but might have slightly different names.
+Wall Street Journal news is drastically different than their opinion section.
+Other organizations have slightly different names for the same thing and a product of the aggregation service and not due to any real difference.
+Luckily, most of the anaylsis is operating on the content of the news headlines, which do not suffer from this data impurity.
+
+\section{Classification Task}
+
+The classification of news titles into emotional categories was accomplished by using a pretrained large langauge model from \href{https://huggingface.co/arpanghoshal/EmoRoBERTa}{HuggingFace}.
+This model was trained on \href{https://ai.googleblog.com/2021/10/goemotions-dataset-for-fine-grained.html}{a dataset curated and published by Google} which manually classified a collection of 58,000 comments into 28 emotions.
+The classes for each article will be derived by tokenizing the title and running the model over the tokens, then grabbing the largest probabilty class from the output.
+
+The data has been discretized into years.
+ Additionally, the publishers will have been discretized based of either principle component analysis on link similarity or based on the bias ratings of \href{https://www.allsides.com/media-bias/ratings}{All Sides}.
+Given that the features of the dataset are sparse, it is not expected to have any useless attributes, unless the original hypothesis of a temporal trend proving to be false.
+Of the features used in the analysis, there are enough data points that null or missing values can safely be excluded.
+
+\section{Experiments}
+
+No computational experiment have been done yet.
+Generating the tokenized text, the word embedding and the emotional sentiment analysis have made up the bulk of the work thus far.
+The bias ratings do not cover all publisher in the dataset, so the number of articles without a bias rating from their publisher will have to be calculated.
+If it is less than 30\% of the articles, it might not make sense to use the bias ratings.
+The creation and reduction of the link graph with principle component analysis will need to be done to visualize the relationship between related publishers.
+
+\section{Results}
+\textbf{TODO.}
+
+\end{multicols}
+
+\end{document}
diff --git a/docs/presentation.md b/docs/presentation.md
new file mode 100644
index 0000000..c6ad8f1
--- /dev/null
+++ b/docs/presentation.md
@@ -0,0 +1,552 @@
+_model: slides
+---
+
+title: CSCI 577 - Data Mining
+
+---
+body:
+
+# Political Polarization
+
+Matt Jensen
+
+===
+
+# Hypothesis
+
+Political polarization is rising, and news articles are a proxy measure.
+
+==
+
+# Is this reasonable?
+
+
+==
+
+# Why is polarization rising?
+
+Not my job, but there's research[ref](#references) to support it
+
+
+==
+
+# Sub-hypothesis
+
+- The polarization increases near elections.
+- The polarization is not evenly distributed across publishers.
+- The polarization is not evenly distributed across political specturm.
+
+==
+
+# Sub-sub-hypothesis
+
+- Similarly polarized publishers link to each other.
+- 'Mainstream' media uses more neutral titles.
+- Highly polarized publications don't last as long.
+
+===
+
+# Data Source(s)
+
+memeorandum.com
+
+allsides.com
+
+huggingface.com
+
+===
+
+
+
+===
+
+# memeorandum.com
+
+- News aggregation site.
+- Was really famous before Google News.
+- Still aggregates sites today.
+
+==
+
+# Why Memeorandum?
+
+- Behavioral: I only read titles sometimes. (doom scrolling).
+- Behavioral: It's my source of news (with sister site TechMeme.com).
+- Convenient: most publishers block bots.
+- Convenient: dead simple html to parse.
+- Archival: all headlines from 2006 forward.
+- Archival: automated, not editorialized.
+
+===
+
+
+
+===
+
+# AllSides.com
+
+- Rates news publications as left, center or right.
+- Ratings combine:
+ - blind bias surveys.
+ - editorial reviews.
+ - third party research.
+ - community voting.
+- Originally scraped website, but direct access eventually.
+
+
+==
+
+# Why AllSides?
+
+- Behavioral: One of the first google results on bias apis.
+- Convenient: Ordinal ratings [-2: very left, 2: very right].
+- Convenient: Easy format.
+- Archival: Covers 1400 publishers.
+
+===
+
+
+
+===
+
+# HuggingFace.com
+
+- Deep Learning library.
+- Lots of pretrained models.
+- Easy, off the shelf word/sentence embeddings and text classification models.
+
+==
+
+# Why HuggingFace?
+
+- Behavioral: Language Models are HOT right now.
+- Behavioral: The dataset needed more features.
+- Convenient: Literally 5 lines of python.
+- Convenient: Testing different model performance was easy.
+- Archival: Lots of pretrained classification tasks.
+
+===
+
+# Data Structures
+Stories
+
+- Top level stories.
+ - title.
+ - publisher.
+ - author.
+- Related discussion.
+ - publisher.
+ - uses 'parent' story as a source.
+- Stream of stories (changes constantly).
+
+==
+
+# Data Structures
+Bias
+
+- Per publisher.
+ - name.
+ - label.
+ - agree/disagree vote by community.
+- Name could be semi-automatically joined to stories.
+
+==
+
+# Data Structures
+Embeddings
+
+- Per story title.
+ - sentence embedding (n, 384).
+ - sentiment classification (n, 1).
+ - emotional classification (n, 1).
+- ~ 1 hour of inference time to map story titles and descriptions.
+
+===
+
+# Data Collection
+
+==
+
+# Data Collection
+
+Story Scraper (simplified)
+
+```python
+day = timedelta(days=1)
+cur = date(2005, 10, 1)
+end = date.today()
+while cur <= end:
+ cur = cur + day
+ save_as = output_dir / f"{cur.strftime('%y-%m-%d')}.html"
+ url = f"https://www.memeorandum.com/{cur.strftime('%y%m%d')}/h2000"
+ r = requests.get(url)
+ with open(save_as, 'w') as f:
+ f.write(r.text)
+```
+
+==
+
+# Data Collection
+Bias Scraper (hard)
+
+```python
+...
+bias_html = DATA_DIR / 'allsides.html'
+parser = etree.HTMLParser()
+tree = etree.parse(str(bias_html), parser)
+root = tree.getroot()
+rows = root.xpath('//table[contains(@class,"views-table")]/tbody/tr')
+
+ratings = []
+for row in rows:
+ rating = dict()
+ ...
+```
+
+==
+
+# Data Collection
+Bias Scraper (easy)
+
+
+
+==
+
+# Data Collection
+Embeddings (easy)
+
+```python
+# table = ...
+tokenizer = AutoTokenizer.from_pretrained("roberta-base")
+model = AutoModel.from_pretrained("roberta-base")
+
+for chunk in table:
+ tokens = tokenizer(chunk, add_special_tokens = True, truncation = True, padding = "max_length", max_length=92, return_attention_mask = True, return_tensors = "pt")
+ outputs = model(**tokens)
+ embeddings = outputs.last_hidden_state.detach().numpy()
+ ...
+```
+
+==
+
+# Data Collection
+Classification Embeddings (medium)
+
+```python
+...
+outputs = model(**tokens)[0].detach().numpy()
+scores = 1 / (1 + np.exp(-outputs)) # Sigmoid
+class_ids = np.argmax(scores, axis=1)
+for i, class_id in enumerate(class_ids):
+ results.append({"story_id": ids[i], "label" : model.config.id2label[class_id]})
+...
+```
+
+===
+
+# Data Selection
+
+==
+
+# Data Selection
+Stories
+
+- Clip the first and last full year of stories.
+- Remove duplicate stories (big stories span multiple days).
+
+==
+# Data Selection
+
+Publishers
+
+- Combine subdomains of stories.
+ - blog.washingtonpost.com and washingtonpost.com are considered the same publisher.
+ - This could be bad. For example: opinion.wsj.com != wsj.com.
+
+==
+
+# Data Selection
+
+Links
+
+- Select only stories with publishers whose story had been a 'parent' ('original publishers').
+ - Eliminates small blogs and non-original news.
+- Eliminate publishers without links to original publishers.
+ - Eliminate silo'ed publications.
+ - Link matrix is square and low'ish dimensional.
+
+==
+
+# Data Selection
+
+Bias
+
+- Keep all ratings, even ones with low agree/disagree ratio.
+- Join datasets on publisher name.
+ - Not automatic (look up Named Entity Recognition).
+ - Started with 'jaro winkler similarity' then manually from there.
+- Use numeric values
+ - [left: -2, left-center: -1, ...]
+
+===
+
+# Descriptive Stats
+
+Raw
+
+| metric | value |
+|:------------------|--------:|
+| total stories | 299714 |
+| total related | 960111 |
+| publishers | 7031 |
+| authors | 34346 |
+| max year | 2023 |
+| min year | 2005 |
+| top level domains | 7063 |
+
+==
+# Descriptive Stats
+
+Stories Per Publisher
+
+
+
+==
+
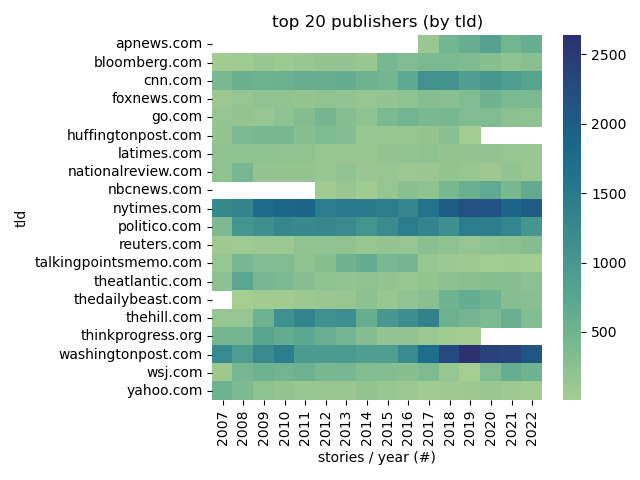
+# Descriptive Stats
+
+Top Publishers
+
+
+
+==
+
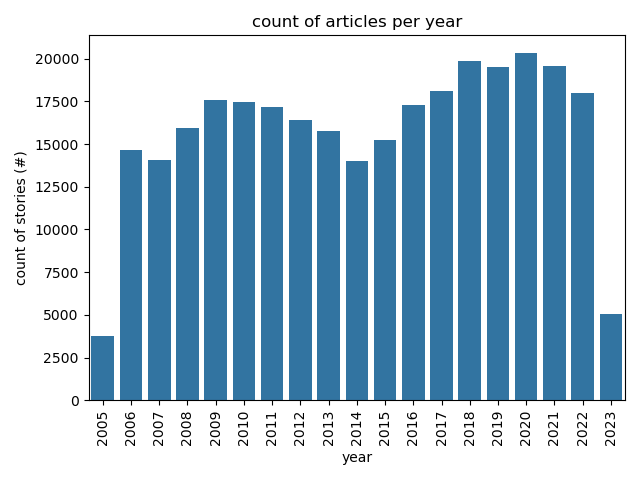
+# Descriptive Stats
+
+Articles Per Year
+
+
+
+==
+
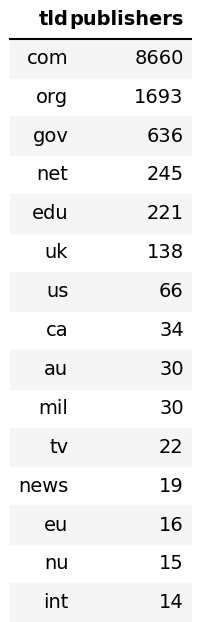
+# Descriptive Stats
+
+Common TLDs
+
+
+
+==
+
+# Descriptive Stats
+
+Post Process
+
+| key | value |
+|:------------------|--------:|
+| total stories | 251553 |
+| total related | 815183 |
+| publishers | 223 |
+| authors | 23809 |
+| max year | 2022 |
+| min year | 2006 |
+| top level domains | 234 |
+
+===
+# Experiments
+
+1. **clustering** on link similarity.
+2. **classification** on link similarity.
+3. **classification** on sentence embedding.
+4. **classification** on sentiment analysis.
+5. **regression** on emotional classification over time and publication.
+
+===
+# Experiment 1
+
+Setup
+
+- Create one-hot encoding of links between publishers.
+- Cluster the encoding.
+- Expect similar publications in same cluster.
+- Use PCA to visualize clusters.
+
+Note:
+Principle Component Analysis:
+- a statistical technique for reducing the dimensionality of a dataset.
+- linear transformation into a new coordinate system where (most of) the variation data can be described with fewer dimensions than the initial data.
+
+==
+
+# Experiment 1
+
+One Hot Encoding
+
+| publisher | nytimes| wsj| newsweek| ...|
+|:----------|--------:|----:|--------:|----:|
+| nytimes | 1| 1| 1| ...|
+| wsj | 1| 1| 0| ...|
+| newsweek | 0| 0| 1| ...|
+| ... | ...| ...| ...| ...|
+
+==
+
+# Experiment 1
+
+n-Hot Encoding
+
+| publisher | nytimes| wsj| newsweek| ...|
+|:----------|--------:|----:|--------:|----:|
+| nytimes | 11| 1| 141| ...|
+| wsj | 1| 31| 0| ...|
+| newsweek | 0| 0| 1| ...|
+| ... | ...| ...| ...| ...|
+
+==
+
+# Experiment 1
+
+Normalized n-Hot Encoding
+
+| publisher | nytimes| wsj| newsweek| ...|
+|:----------|--------:|----:|--------:|----:|
+| nytimes | 0| 0.4| 0.2| ...|
+| wsj | 0.2| 0| 0.4| ...|
+| newsweek | 0.0| 0.0| 0.0| ...|
+| ... | ...| ...| ...| ...|
+
+==
+
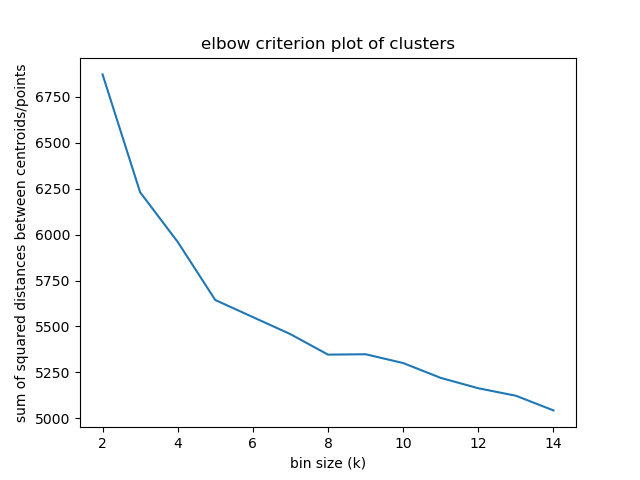
+# Experiment 1
+
+Elbow criterion
+
+
+
+Note:
+
+The elbow method looks at the percentage of explained variance as a function of the number of clusters:
+
+One should choose a number of clusters so that adding another cluster doesn't give much better modeling of the data.
+
+Percentage of variance explained is the ratio of the between-group variance to the total variance,
+
+==
+
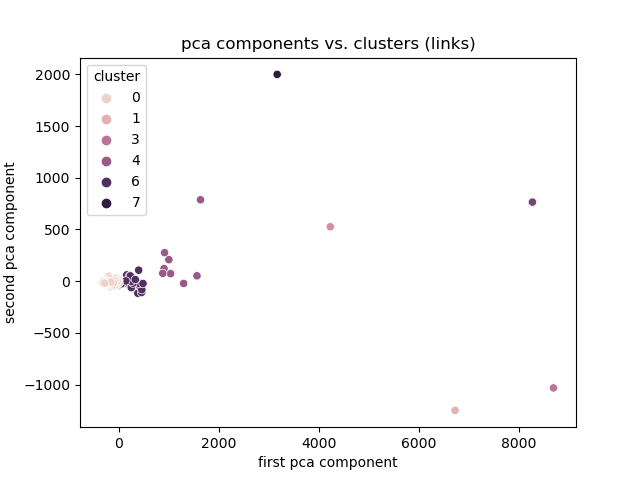
+# Experiment 1
+
+Link Magnitude
+
+
+
+==
+
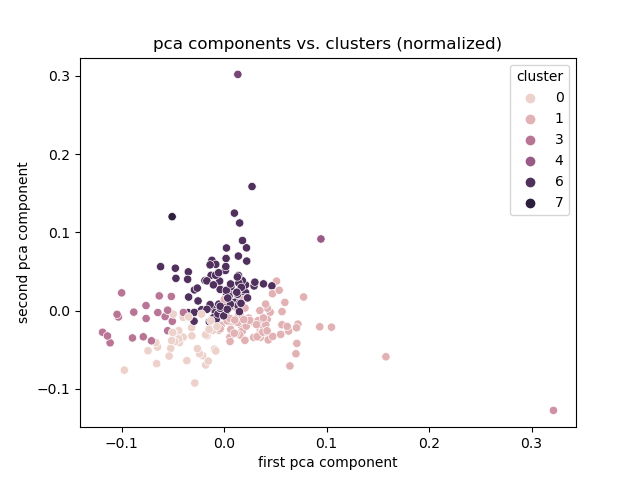
+# Experiment 1
+
+Normalized
+
+
+
+==
+
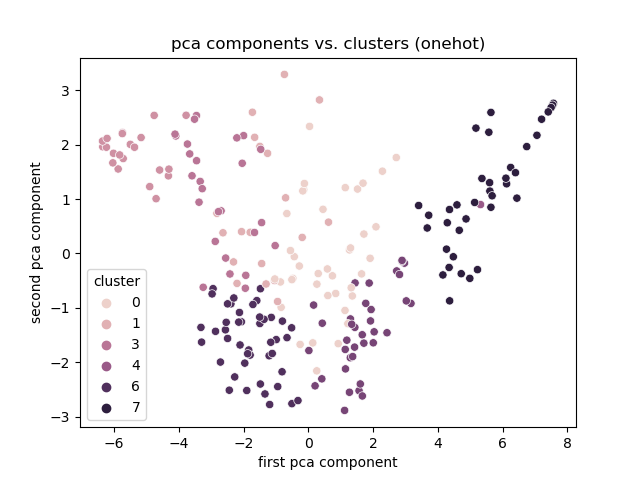
+# Experiment 1
+
+Onehot
+
+
+
+==
+
+# Experiment 1
+
+Discussion
+
+- Best encoding: One hot.
+ - Clusters based on total links otherwise.
+- Clusters, but no explanation
+- Limitation: need the link encoding to cluster.
+ - Smaller publishers might not link very much.
+
+===
+
+# Experiment 2
+
+Setup
+
+- Create features. :
+ - Publisher frequency.
+ - Reuse link encodings.
+- Create classes:
+ - Join bias classifications.
+- Train classifier.
+
+Note:
+
+==
+# Experiment 2
+Descriptive stats
+
+| metric | value |
+|:------------|:----------|
+| publishers | 1582 |
+| labels | 6 |
+| left | 482 |
+| center | 711 |
+| right | 369 |
+| agree range | [0.0-1.0] |
+
+==
+
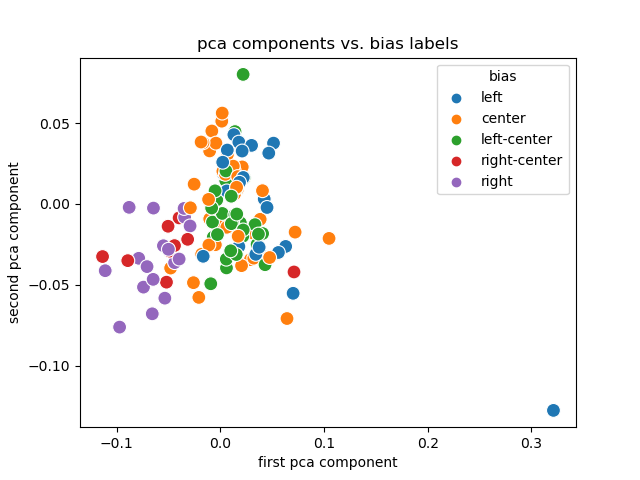
+# Experiment 2
+
+PCA + Labels
+
+
+
+==
+
+# Experiment 2
+
+Discussion
+
+- Link encodings (and their PCA) are useful.
+ - Labels are (sort of) separated and clustered.
+ - Creating them for smaller publishers is trivial.
+==
+
+# Experiment 2
+
+Limitations
+
+- Dependent on accurate rating.
+- Ordinal ratings not available.
+- Dependent on accurate joining across datasets.
+- Entire publication is rated, not authors.
+- Don't know what to do with community rating.
+
+===
+
+# Experiment 3
+
+Setup
+
+==
+
+# Limitations
+
+- Many different authors under the same publisher.
+- Publishers use syndication.
+- Bias ratings are biased.
+
+===
+
+# Questions
+
+===
+
+
+
+# References
+
+[1]: Stewart, A.J. et al. 2020. Polarization under rising inequality and economic decline. Science Advances. 6, 50 (Dec. 2020), eabd4201. DOI:https://doi.org/10.1126/sciadv.abd4201.
+
+Note:
diff --git a/src/bias.py b/src/bias.py
index 5347aa7..92f952a 100644
--- a/src/bias.py
+++ b/src/bias.py
@@ -1,12 +1,12 @@
import click
-from data import connect
+from data.main import connect
import pandas as pd
from lxml import etree
from pathlib import Path
import os
import csv
-def map(rating:str) -> int:
+def label_to_int(rating:str) -> int:
mapping = {
'left' : 0,
@@ -19,20 +19,18 @@ def map(rating:str) -> int:
return mapping[rating]
+def int_to_label(class_id: int) -> str:
+ mapping = {
+ 0 : 'left',
+ 1 : 'left-center',
+ 2 : 'center',
+ 3 : 'right-center',
+ 4 : 'right',
+ -1 : 'allsides',
+ }
+ return mapping[class_id]
-@click.command(name="bias:load")
-def load() -> None:
- DB = connect()
- DATA_DIR = Path(os.environ['DATA_MINING_DATA_DIR'])
- f = str(DATA_DIR / "bias_ratings.csv")
- DB.sql(f"""
- create table bias_ratings as
- select
- row_number() over(order by b.publisher) as id
- ,b.*
- from read_csv_auto('{f}') b
- """)
@click.command(name="bias:normalize")
def normalize() -> None:
DB = connect()
@@ -41,133 +39,48 @@ def normalize() -> None:
CREATE OR REPLACE TABLE publisher_bias AS
WITH cte AS (
SELECT
- p.id
+ p.id as publisher_id
+ ,b.id as bias_id
,b.bias as label
,JARO_WINKLER_SIMILARITY(LOWER(p.name), LOWER(b.publisher)) as similarity
FROM bias_ratings b
- JOIN publishers p
+ JOIN top.publishers p
ON JARO_WINKLER_SIMILARITY(LOWER(p.name), LOWER(b.publisher)) > 0.95
),ranked AS (
SELECT
- id
+ publisher_id
+ ,bias_id
,label
,similarity
- ,ROW_NUMBER() OVER(PARTITION BY id ORDER BY similarity DESC) AS rn
+ ,ROW_NUMBER() OVER(PARTITION BY publisher_id ORDER BY similarity DESC) AS rn
FROM cte
)
SELECT
- id
+ publisher_id
,label
+ ,bias_id
FROM ranked
WHERE ranked.rn = 1
""")
+ mapping = [
+ {'label' :'left' , 'ordinal': -2},
+ {'label' :'left-center' , 'ordinal': -1},
+ {'label' :'center' , 'ordinal': 0},
+ {'label' :'right-center' , 'ordinal': 1},
+ {'label' :'right' , 'ordinal': 2},
+ ]
+ mapping = pd.DataFrame(mapping)
- DB.sql("""
- with cte as (
- select
- s.publisher_id
- ,count(1) as stories
- from stories s
- group by s.publisher_id
- )
- select
- s.publisher
- ,s.stories
- ,b.publisher
- ,b.bias
- from bias_ratings b
- join cte s
- on s.publisher = b.publisher
- order by
- stories desc
- limit 15
+ DB.query("alter table bias_ratings add column ordinal int")
+
+ DB.query("""
+ update bias_ratings b
+ set ordinal = o.ordinal
+ FROM mapping o
+ WHERE o.label = b.bias
""")
- DB.sql("""
- with cte as (
- select
- s.publisher
- ,count(1) as stories
- from stories s
- group by s.publisher
- )
- select
- sum(stories)
- ,avg(agree / disagree)
- from bias_ratings b
- join cte s
- on s.publisher = b.publisher
- """)
-
- DB.sql("""
- with cte as (
- select
- s.publisher
- ,count(1) as stories
- from stories s
- group by s.publisher
- )
- select
- sum(s.stories) filter(where b.publisher is not null) as matched
- ,sum(s.stories) filter(where b.publisher is null) as unmatched
- ,cast(sum(s.stories) filter(where b.publisher is not null) as numeric)
- / sum(s.stories) filter(where b.publisher is null) as precent_matched
- from bias_ratings b
- right join cte s
- on s.publisher = b.publisher
- """)
-
- DB.sql("""
- select
- *
- from bias_ratings
- where publisher ilike '%CNN%'
- """)
-
-@click.command(name='bias:debug')
-def debug() -> None:
- DB = connect()
- DATA_DIR = Path(os.environ['DATA_MINING_DATA_DIR'])
- f = str(DATA_DIR / "bias_ratings.csv")
-
- DB.sql("""
- with cte as (
- select
- outlet
- ,count(1) as stories
- from stories
- group by outlet
- )
- ,total as (
- select
- sum(stories) as total
- from cte
- )
- select
- cte.outlet
- ,cte.stories
- ,bias.outlet
- ,bias.lean
- ,sum(100 * (cte.stories / cast(total.total as float))) over() as rep
- ,total.total
- from cte
- join bias
- on jaro_winkler_similarity(bias.outlet, cte.outlet) > 0.9
- cross join total.total
- """)
-
- DB.sql("""
- select
- outlet
- ,count(1) as stories
- from stories
- group by outlet
- order by count(1) desc
- limit 50
- """)
-
- outlets
@click.command(name='bias:parse')
def parse() -> None:
@@ -199,3 +112,64 @@ def parse() -> None:
ratings.append(rating)
df = pd.DataFrame(ratings)
df.to_csv(DATA_DIR / 'bias_ratings.csv', sep="|", index=False, quoting=csv.QUOTE_NONNUMERIC)
+
+@click.command(name="bias:load")
+def load() -> None:
+ DB = connect()
+ DATA_DIR = Path(os.environ['DATA_MINING_DATA_DIR'])
+ f = str(DATA_DIR / "bias_ratings.csv")
+
+ DB.sql(f"""
+ CREATE TABLE bias_ratings as
+ select
+ row_number() over(order by b.publisher) as id
+ ,b.*
+ from read_csv_auto('{f}') b
+ """)
+
+@click.command('bias:export')
+def export():
+ data_path = Path(os.environ['DATA_MINING_DATA_DIR'])
+
+ DB = connect()
+ all_bias = DB.query("""
+ SELECT
+ id as bias_id
+ ,publisher as name
+ ,bias as label
+ FROM bias_ratings
+ ORDER by agree desc
+ """)
+ all_bias.df().to_csv(data_path / 'TMP_publisher_bias.csv', sep="|", index=False)
+ mapped_bias = DB.query("""
+ SELECT
+ p.id as publisher_id
+ ,p.name as name
+ ,p.tld as tld
+ ,b.label as bias
+ ,b.bias_id as bias_id
+ FROM top.publishers p
+ LEFT JOIN publisher_bias b
+ ON b.publisher_id = p.id
+ """)
+ mapped_bias.df().to_csv(data_path / 'TMP_publisher_bias_to_load.csv', sep="|", index=False)
+ DB.close()
+
+@click.command('bias:import-mapped')
+def import_mapped():
+ data_path = Path(os.environ['DATA_MINING_DATA_DIR'])
+ table_name = "top.publisher_bias"
+
+ DB = connect()
+ df = pd.read_csv(data_path / 'TMP_publisher_bias_to_load.csv', sep="|")
+
+ DB.query(f"""
+ CREATE OR REPLACE TABLE {table_name} AS
+ SELECT
+ publisher_id AS publisher_id
+ ,cast(bias_id AS int) as bias_id
+ FROM df
+ WHERE bias_id IS NOT NULL
+ """)
+ print(f"created table: {table_name}")
+
diff --git a/src/cli.py b/src/cli.py
index d91e0e2..11744d3 100644
--- a/src/cli.py
+++ b/src/cli.py
@@ -7,7 +7,7 @@ def cli():
if __name__ == "__main__":
load_dotenv()
- import scrape
+ from data import scrape
cli.add_command(scrape.download)
cli.add_command(scrape.parse)
cli.add_command(scrape.load)
@@ -32,4 +32,26 @@ if __name__ == "__main__":
cli.add_command(emotion.create_table)
import sentence
cli.add_command(sentence.embed)
+ from train import main as train_main
+ cli.add_command(train_main.main)
+
+ import plots.descriptive as plotd
+ cli.add_command(plotd.articles_per_year)
+ cli.add_command(plotd.distinct_publishers)
+ cli.add_command(plotd.stories_per_publisher)
+ cli.add_command(plotd.top_publishers)
+ cli.add_command(plotd.common_tld)
+
+ import links as linkcli
+ cli.add_command(linkcli.create_table)
+ cli.add_command(linkcli.create_pca)
+ cli.add_command(linkcli.create_clusters)
+
+ import plots.links as plotl
+ cli.add_command(plotl.elbow)
+ cli.add_command(plotl.link_pca_clusters)
+
+ import plots.classifier as plotc
+ cli.add_command(plotc.pca_with_classes)
+
cli()
diff --git a/src/data/__init__.py b/src/data/__init__.py
new file mode 100644
index 0000000..0c64a7f
--- /dev/null
+++ b/src/data/__init__.py
@@ -0,0 +1,6 @@
+import data.main
+import data.scrape
+__all__ = [
+ 'main'
+ ,'scrape'
+]
diff --git a/src/data.py b/src/data/main.py
similarity index 100%
rename from src/data.py
rename to src/data/main.py
diff --git a/src/scrape.py b/src/data/scrape.py
similarity index 73%
rename from src/scrape.py
rename to src/data/scrape.py
index f3a285c..13377ba 100644
--- a/src/scrape.py
+++ b/src/data/scrape.py
@@ -4,10 +4,12 @@ import requests
from pathlib import Path
import click
from tqdm import tqdm
-from data import data_dir, connect
+from data.main import data_dir, connect
from lxml import etree
import pandas as pd
from urllib.parse import urlparse
+from tld import get_tld
+from tld.utils import update_tld_names
@click.command(name='scrape:load')
@click.option('--directory', type=Path, default=data_dir(), show_default=True)
@@ -61,6 +63,7 @@ def download(output_dir):
@click.option('-o', '--output_dir', type=Path, default=data_dir(), show_default=True)
def parse(directory, output_dir):
"""parse the html files on disk into a structured csv format."""
+ update_tld_names()
directory = data_dir() / "memeorandum"
parser = etree.HTMLParser()
pages = [f for f in directory.glob("*.html")]
@@ -104,8 +107,7 @@ def parse(directory, output_dir):
url = item.xpath('.//strong/a')[0].get('href')
out['url'] = url
- out['publisher_url_domain'] = urlparse(publisher_url).netloc
- out['domain'] = urlparse(url).netloc
+ out['tld'] = get_tld(publisher_url)
item_id = hash((page.stem, url))
out['id'] = item_id
@@ -225,3 +227,111 @@ def normalize():
alter table related_stories drop publisher_domain;
""")
+
+def another_norm():
+ sv2 = pd.read_csv(data_dir / 'stories.csv', sep="|")
+ related = pd.read_csv(data_dir / 'related.csv', sep="|")
+
+ related['tld'] = related.url.apply(lambda x: map_tld(x))
+
+ DB.query("""
+ update related_stories
+ set publisher_id = p.id
+ from publishers p
+ join related r
+ on r.tld = p.tld
+ where r.url = related_stories.url
+ """)
+
+
+ DB.query("""alter table stories add column tld text""")
+
+ s_url = DB.query("""
+ select
+ id
+ ,url
+ from stories
+ """).df()
+
+
+ s_url['tld'] = s_url.url.apply(lambda x: map_tld(x))
+
+ DB.query("""
+ update stories
+ set tld = s_url.tld
+ from s_url
+ where s_url.id = stories.id
+ """)
+
+ DB.query("""
+ update stories
+ set publisher_id = p.id
+ from publishers p
+ where p.tld = stories.tld
+ """)
+
+
+ select
+ DB.query("""
+ update stories
+ set stories.publisher_id = p.id
+ from new_pub
+ """)
+ sv2['tld'] = sv2.publisher_url.apply(lambda x: map_tld(x))
+
+
+ new_pub = DB.query("""
+ with cte as (
+ select
+ tld
+ ,publisher
+ ,count(1) filter(where year(published_at) = 2022) as recent_ctn
+ ,count(1) as ctn
+ from sv2
+ group by
+ tld
+ ,publisher
+ )
+ ,r as (
+ select
+ tld
+ ,publisher
+ ,ctn
+ ,row_number() over(partition by tld order by recent_ctn desc) as rn
+ from cte
+ )
+ select
+ row_number() over() as id
+ ,publisher as name
+ ,tld
+ from r
+ where rn = 1
+ order by ctn desc
+ """).df()
+
+ DB.query("""
+ CREATE OR REPLACE TABLE publishers AS
+ SELECT
+ id
+ ,name
+ ,tld
+ FROM new_pub
+ """)
+
+
+ def map_tld(x):
+ try:
+ res = get_tld(x, as_object=True)
+ return res.fld
+ except:
+ return None
+
+ DB.sql("""
+ SELECT
+ s.id
+ ,sv2.publisher_url
+ FROM stories s
+ JOIN sv2
+ on sv2.id = s.id
+ limit 5
+ """)
diff --git a/src/emotion.py b/src/emotion.py
index 7ef23e6..b7e4824 100644
--- a/src/emotion.py
+++ b/src/emotion.py
@@ -6,7 +6,7 @@ import numpy as np
from transformers import BertTokenizer
from model import BertForMultiLabelClassification
-from data import connect, data_dir
+from data.main import connect, data_dir
import seaborn as sns
import matplotlib.pyplot as plt
from matplotlib.dates import DateFormatter
@@ -376,3 +376,99 @@ def debug():
DB.close()
out.to_csv(data_dir() / 'emotions.csv', sep="|")
+
+def another():
+ DB = connect()
+ DB.sql("""
+ select
+ *
+ from emotions
+ """)
+
+ emotions = DB.sql("""
+ select
+ year(s.published_at) as year
+ ,se.label as emotion
+ ,count(1) as stories
+ from stories s
+ join story_emotions se
+ on s.id = se.story_id
+ group by
+ year(s.published_at)
+ ,se.label
+ """).df()
+
+ sns.scatterplot(x=emotions['year'], y=emotions['stories'], hue=emotions['emotion'])
+ plt.show()
+
+ pivot = emotions.pivot(index='year', columns='emotion', values='stories')
+ pivot.reset_index(inplace=True)
+ from sklearn.linear_model import LinearRegression
+ reg = LinearRegression()
+
+ for emotion in pivot.keys()[1:].tolist():
+ _ = reg.fit(pivot['year'].to_numpy().reshape(-1, 1), pivot[emotion])
+ print(f"{emotion}: {reg.coef_[0]}")
+
+ fig, ax = plt.subplots()
+ #sns.lineplot(x=pivot['anger'], y=pivot['joy'])
+ #sns.lineplot(x=pivot['anger'], y=pivot['surprise'], ax=ax)
+ sns.lineplot(x=pivot['anger'], y=pivot['fear'], ax=ax)
+ sns.lineplot(x=pivot[''], y=pivot['fear'], ax=ax)
+ plt.show()
+
+ DB.close()
+
+ normalized = DB.sql("""
+ with cte as (
+ select
+ year(s.published_at) as year
+ ,se.label as emotion
+ ,b.label as bias
+ from stories s
+ join story_emotions se
+ on s.id = se.story_id
+ join publisher_bias b
+ on b.id = s.publisher_id
+ where b.label != 'allsides'
+ and se.label != 'neutral'
+ )
+ select
+ distinct
+ year
+ ,emotion
+ ,bias
+ ,cast(count(1) over(partition by year, bias, emotion) as float) / count(1) over(partition by year, bias) as group_count
+ from cte
+ """).df()
+
+ DB.sql("""
+ select
+ b.label as bias
+ ,count(1) as stories
+ from stories s
+ join story_emotions se
+ on s.id = se.story_id
+ join publisher_bias b
+ on b.id = s.publisher_id
+ group by
+ b.label
+ """).df()
+
+ another_pivot = emotional_bias.pivot(index=['bias', 'year'], columns='emotion', values='stories')
+ another_pivot.reset_index(inplace=True)
+
+ sns.lineplot(data=normalized, x='year', y='group_count', hue='bias', style='emotion')
+ plt.show()
+
+ sns.relplot(
+ data=normalized, x="year", y="group_count", hue="emotion", col='bias', kind="line"
+ #data=normalized, x="year", y="group_count", hue="emotion", col='bias', kind="line", facet_kws=dict(sharey=False)
+ )
+ plt.show()
+
+ DB.sql("""
+ select
+ *
+ from another_pivot
+ """)
diff --git a/src/lib.py b/src/lib.py
deleted file mode 100644
index deef1b2..0000000
--- a/src/lib.py
+++ /dev/null
@@ -1,8 +0,0 @@
-import sklearn
-import polars as pl
-import toml
-from pathlib import Path
-
-config = toml.load('/home/user/577/repo/config.toml')
-app_dir = Path(config.get('app').get('path'))
-df = pl.read_csv(app_dir / "data/articles.csv")
diff --git a/src/links.py b/src/links.py
index 93e7cfa..9dc2a56 100644
--- a/src/links.py
+++ b/src/links.py
@@ -1,12 +1,148 @@
-from data import connect
+import click
+from data.main import connect
import pandas as pd
import numpy as np
-from sklearn.decomposition import PCA, TruncatedSVD
-from sklearn.cluster import MiniBatchKMeans
import seaborn as sns
import matplotlib.pyplot as plt
+@click.command('links:create-table')
+def create_table():
+
+ table_name = "top.link_edges"
+ DB = connect()
+ DB.query(f"""
+ CREATE OR REPLACE TABLE {table_name} AS
+ with cte as(
+ SELECT
+ s.publisher_id as parent_id
+ ,r.publisher_id as child_id
+ ,count(1) as links
+ FROM top.stories s
+ JOIN top.related_stories r
+ ON s.id = r.parent_id
+ group by
+ s.publisher_id
+ ,r.publisher_id
+ )
+ SELECT
+ cte.parent_id
+ ,cte.child_id
+ ,cte.links as links
+ ,cast(cte.links as float) / sum(cte.links) over(partition by cte.parent_id) as normalized
+ ,case when cte.links > 0 then 1 else 0 end as onehot
+ FROM cte
+ WHERE cte.child_id in (
+ SELECT
+ distinct parent_id
+ FROM cte
+ )
+ AND cte.parent_id in (
+ SELECT
+ distinct child_id
+ FROM cte
+ )
+ """)
+ DB.close()
+
+ DB = connect()
+ DB.query("""
+ SELECT
+ *
+ ,-log10(links)
+ --distinct parent_id
+ FROM top.link_edges e
+ WHERE e.parent_id = 238
+ """)
+ DB.close()
+ print(f"created {table_name}")
+
+@click.command('links:create-pca')
+@click.option('--source', type=click.Choice(['links', 'normalized', 'onehot']), default='links')
+def create_pca(source):
+ """create 2D pca labels"""
+
+ from sklearn.decomposition import PCA
+
+ table_name = f"top.publisher_pca_{source}"
+ DB = connect()
+ pub = DB.query("""
+ SELECT
+ *
+ FROM top.publishers
+ """).df()
+ df = DB.query(f"""
+ SELECT
+ parent_id
+ ,child_id
+ ,{source} as links
+ FROM top.link_edges
+ """).df()
+ DB.close()
+ pivot = df.pivot(index='parent_id', columns='child_id', values='links').fillna(0)
+
+ svd = PCA(n_components=2)
+ svd_out = svd.fit_transform(pivot)
+
+ out = pivot.reset_index()[['parent_id']]
+ out['first'] = svd_out[:, 0]
+ out['second'] = svd_out[:, 1]
+ out = pd.merge(out, pub, left_on='parent_id', right_on='id')
+
+ DB = connect()
+ DB.query(f"""
+ CREATE OR REPLACE TABLE {table_name} AS
+ SELECT
+ out.id as publisher_id
+ ,out.first as first
+ ,out.second as second
+ FROM out
+ """)
+ DB.close()
+ print(f"created {table_name}")
+
+
+@click.command('links:create-clusters')
+@click.option('--source', type=click.Choice(['links', 'normalized', 'onehot']), default='links')
+def create_clusters(source):
+ from sklearn.cluster import KMeans
+
+ table_name = f"top.publisher_clusters_{source}"
+ DB = connect()
+ df = DB.query(f"""
+ SELECT
+ parent_id
+ ,child_id
+ ,{source} as links
+ FROM top.link_edges
+ """).df()
+ pub = DB.query("""
+ SELECT
+ *
+ FROM top.publishers
+ """).df()
+ DB.close()
+ pivot = df.pivot(index='parent_id', columns='child_id', values='links').fillna(0)
+
+
+ k = 8
+ kmeans = KMeans(n_clusters=k, n_init="auto")
+ pred = kmeans.fit_predict(pivot)
+ out = pivot.reset_index()[['parent_id']]
+ out['label'] = pred
+ out = pd.merge(out, pub, left_on='parent_id', right_on='id')
+ new_table = out[['id', 'label']]
+
+ DB = connect()
+ DB.query(f"""
+ CREATE OR REPLACE TABLE {table_name} AS
+ SELECT
+ n.id as publisher_id
+ ,n.label as label
+ FROM new_table n
+ """)
+ DB.close()
+ print(f"created {table_name}")
def to_matrix():
"""returns an adjacency matrix of publishers to publisher link frequency"""
@@ -21,6 +157,7 @@ def to_matrix():
{'label' :'right', 'value' : 4},
{'label' :'allsides', 'value' : -1},
])
+
bias = DB.sql("""
SELECT
b.id
@@ -37,11 +174,7 @@ def to_matrix():
p.id
,p.name
,p.url
- ,b.label
- ,b.value
from publishers p
- left join bias b
- on b.id = p.id
""").df()
edges = DB.sql("""
@@ -81,12 +214,23 @@ def to_matrix():
ON p.id = cte.parent_id
""").df()
+ # only keep values that have more than 1 link
+ test = edges[edges['links'] > 2].pivot(index='parent_id', columns='child_id', values='links').fillna(0).reset_index()
+ edges.dropna().pivot(index='parent_id', columns='child_id', values='links').fillna(0)
+ pd.merge(adj, pub, how='left', left_on='parent_id', right_on='id')
adj = edges.pivot(index='parent_id', columns='child_id', values='links').fillna(0)
+ adj.values.shape
out = pd.DataFrame(adj.index.values, columns=['id'])
out = pd.merge(out, pub, how='left', on='id')
+ return out
+@click.command('links:analysis')
+def analysis():
+ from sklearn.decomposition import PCA, TruncatedSVD
+ from sklearn.cluster import MiniBatchKMeans
+ adj = to_matrix()
pca = PCA(n_components=4)
pca_out = pca.fit_transform(adj)
diff --git a/src/mine.py b/src/mine.py
index d7f1d29..8c2108b 100644
--- a/src/mine.py
+++ b/src/mine.py
@@ -1,4 +1,4 @@
-from data import data_dir, connect
+from data.main import data_dir, connect
import numpy as np
import sklearn
from sklearn.cluster import MiniBatchKMeans
diff --git a/src/plots/__init__.py b/src/plots/__init__.py
new file mode 100644
index 0000000..e69de29
diff --git a/src/plots/classifier.py b/src/plots/classifier.py
new file mode 100644
index 0000000..c85aa7d
--- /dev/null
+++ b/src/plots/classifier.py
@@ -0,0 +1,34 @@
+import click
+from data.main import connect
+import os
+import seaborn as sns
+import matplotlib.pyplot as plt
+from pathlib import Path
+
+out_dir = Path(os.getenv('DATA_MINING_DOC_DIR')) / 'figures'
+
+@click.command('plot:pca-with-classes')
+def pca_with_classes():
+ filename = "pca_with_classes.png"
+
+ DB = connect()
+ data = DB.query(f"""
+ SELECT
+ p.tld
+ ,b.bias
+ ,c.first
+ ,c.second
+ ,round(cast(b.agree as float) / (b.agree + b.disagree), 2) ratio
+ FROM top.publishers p
+ JOIN top.publisher_bias pb
+ ON p.id = pb.publisher_id
+ JOIN bias_ratings b
+ ON b.id = pb.bias_id
+ JOIN top.publisher_pca_normalized c
+ ON c.publisher_id = p.id
+ """).df()
+ DB.close()
+ ax = sns.scatterplot(x=data['first'], y=data['second'], hue=data['bias'], s=100)
+ ax.set(title="pca components vs. bias labels", xlabel="first pca component", ylabel="second pca component")
+ plt.savefig(out_dir / filename)
+ print(f"saved: {filename}")
diff --git a/src/plots/descriptive.py b/src/plots/descriptive.py
new file mode 100644
index 0000000..24cf25b
--- /dev/null
+++ b/src/plots/descriptive.py
@@ -0,0 +1,302 @@
+import click
+from data.main import connect
+import os
+import seaborn as sns
+import matplotlib.pyplot as plt
+from pathlib import Path
+import numpy as np
+
+out_dir = Path(os.getenv('DATA_MINING_DOC_DIR')) / 'figures'
+
+@click.command('plot:articles-per-year')
+def articles_per_year():
+ filename = 'articles_per_year.png'
+
+ DB = connect()
+ data = DB.query("""
+ select
+ year(published_at) as year
+ ,count(1) as stories
+ from stories
+ group by
+ year(published_at)
+ """).df()
+ DB.close()
+
+ ax = sns.barplot(x=data.year, y=data.stories, color='tab:blue')
+ ax.tick_params(axis='x', rotation=90)
+ ax.set(title="count of articles per year", ylabel="count of stories (#)")
+ plt.tight_layout()
+ plt.savefig(out_dir / filename)
+
+@click.command('plot:distinct-publishers')
+def distinct_publishers():
+ filename = 'distinct_publishers.png'
+
+ DB = connect()
+ data = DB.query("""
+ select
+ year(published_at) as year
+ ,count(distinct publisher_id) as publishers
+ from stories
+ group by
+ year(published_at)
+ """).df()
+ DB.close()
+
+ ax = sns.barplot(x=data.year, y=data.publishers, color='tab:blue')
+ ax.tick_params(axis='x', rotation=90)
+ ax.set(title="count of publishers per year", ylabel="count of publishers (#)")
+ plt.tight_layout()
+ plt.savefig(out_dir / filename)
+ plt.close()
+
+@click.command('plot:stories-per-publisher')
+def stories_per_publisher():
+ filename = 'stories_per_publisher.png'
+
+ DB = connect()
+ data = DB.query("""
+ with cte as (
+ select
+ publisher_id
+ ,year(published_at) as year
+ ,count(1) as stories
+ from stories
+ group by
+ publisher_id
+ ,year(published_at)
+ ) , agg as (
+ select
+ publisher_id
+ ,avg(stories) as stories_per_year
+ ,case
+ when avg(stories) < 2 then 2
+ when avg(stories) < 4 then 4
+ when avg(stories) < 8 then 8
+ when avg(stories) < 16 then 16
+ when avg(stories) < 32 then 32
+ when avg(stories) < 64 then 64
+ when avg(stories) < 128 then 128
+ else 129
+ end as max_avg
+ from cte
+ group by
+ publisher_id
+ )
+ select
+ max_avg
+ ,count(1) as publishers
+ from agg
+ group by
+ max_avg
+ """).df()
+ DB.close()
+
+ ax = sns.barplot(x=data.max_avg, y=data.publishers, color='tab:blue')
+ ax.set(title="histogram of publisher stories per year", ylabel="count of publishers (#)", xlabel="max average stories / year")
+ plt.tight_layout()
+ plt.savefig(out_dir / filename)
+ plt.close()
+
+
+@click.command('plot:top-publishers')
+def top_publishers():
+ """plot top publishers over time"""
+
+ filename = 'top_publishers.png'
+
+ DB = connect()
+ data = DB.query("""
+ select
+ p.tld
+ ,year(published_at) as year
+ ,count(1) as stories
+ from (
+ select
+ p.tld
+ ,p.id
+ from top.publishers p
+ join top.stories s
+ on s.publisher_id = p.id
+ group by
+ p.tld
+ ,p.id
+ order by count(1) desc
+ limit 20
+ ) p
+ join top.stories s
+ on s.publisher_id = p.id
+ group by
+ p.tld
+ ,year(published_at)
+ order by count(distinct s.id) desc
+ """).df()
+ DB.close()
+
+ pivot = data.pivot(columns='year', index='tld', values='stories')
+ ax = sns.heatmap(pivot, cmap="crest")
+ ax.set(title="top 20 publishers (by tld)", ylabel="tld", xlabel="stories / year (#)")
+ plt.tight_layout()
+ plt.savefig(out_dir / filename)
+ plt.close()
+
+
+@click.command('plot:common_tld')
+def common_tld():
+ import dataframe_image as dfi
+ filename = 'common_tld.png'
+
+ DB = connect()
+ data = DB.query("""
+ select
+ split_part(url, '.', -1) as tld
+ ,count(1) as publishers
+ ,case when count(1) < 20
+ then string_agg(distinct url, '\t')
+ else NULL
+ end as urls
+ from publishers
+ group by
+ split_part(url, '.', -1)
+ order by
+ count(1) desc
+ """).df()
+ DB.close()
+ data[:15][['tld', 'publishers']].style.hide(axis="index").export_png(out_dir / filename, table_conversion='matplotlib')
+
+def stats():
+
+ # raw
+ DB.query("""
+ SELECT
+ 'total stories' as key
+ ,COUNT(1) as value
+ FROM stories
+ UNION
+ SELECT
+ 'total related' as key
+ ,COUNT(1) as value
+ FROM related_stories
+ UNION
+ SELECT
+ 'top level domains' as key
+ ,COUNT(distinct tld) as value
+ FROM stories
+ UNION
+ SELECT
+ 'publishers' as key
+ ,COUNT(1) as value
+ FROM publishers
+ UNION
+ SELECT
+ 'authors' as key
+ ,COUNT(distinct author) as value
+ FROM stories
+ UNION
+ SELECT
+ 'min year' as key
+ ,min(year(published_at)) as value
+ FROM stories
+ UNION
+ SELECT
+ 'max year' as key
+ ,max(year(published_at)) as value
+ FROM stories
+ """).df().to_markdown(index=False)
+
+ # selected
+ DB.query("""
+ SELECT
+ 'total stories' as key
+ ,COUNT(1) as value
+ FROM top.stories
+ UNION
+ SELECT
+ 'total related' as key
+ ,COUNT(1) as value
+ FROM top.related_stories
+ UNION
+ SELECT
+ 'top level domains' as key
+ ,COUNT(distinct tld) as value
+ FROM top.stories
+ UNION
+ SELECT
+ 'publishers' as key
+ ,COUNT(1) as value
+ FROM top.publishers
+ UNION
+ SELECT
+ 'authors' as key
+ ,COUNT(distinct author) as value
+ FROM top.stories
+ UNION
+ SELECT
+ 'min year' as key
+ ,min(year(published_at)) as value
+ FROM top.stories
+ UNION
+ SELECT
+ 'max year' as key
+ ,max(year(published_at)) as value
+ FROM top.stories
+ """).df().to_markdown(index=False)
+
+@click.command('plot:bias-stats')
+def bias_stats():
+ import dataframe_image as dfi
+ filename = 'bias_stats.png'
+
+ DB = connect()
+
+ df = DB.query("""
+ SELECT
+ string_agg(distinct bias)
+ FROM bias_ratings
+ """).df()
+
+ df.keys()
+
+ df = DB.query("""
+ SELECT
+ 'publishers' as metric
+ ,count(1) as value
+ FROM bias_ratings
+ UNION
+ SELECT
+ 'labels' as metric
+ ,count(distinct bias) as value
+ FROM bias_ratings
+ UNION
+ SELECT
+ 'right' as metric
+ ,count(1) as value
+ FROM bias_ratings
+ WHERE bias in ('right', 'right-center')
+ UNION
+ SELECT
+ 'left' as metric
+ ,count(1) as value
+ FROM bias_ratings
+ WHERE bias in ('left', 'left-center')
+ UNION
+ SELECT
+ 'center' as metric
+ ,count(1) as value
+ FROM bias_ratings
+ WHERE bias in ('center')
+ UNION
+ SELECT
+ 'agree_range' as metric
+ ,'['
+ || min(cast(agree as float) / (agree + disagree))
+ || '-'
+ || max(cast(agree as float) / (agree + disagree))
+ || ']'
+ as value
+ FROM bias_ratings
+ WHERE bias in ('center')
+ """).df()
+ DB.close()
+ print(df.to_markdown(index=False))
diff --git a/src/plots/links.py b/src/plots/links.py
new file mode 100644
index 0000000..6526142
--- /dev/null
+++ b/src/plots/links.py
@@ -0,0 +1,114 @@
+import click
+from data.main import connect
+from links import to_matrix
+import os
+import seaborn as sns
+import matplotlib.pyplot as plt
+from pathlib import Path
+import numpy as np
+from sklearn.metrics import silhouette_score
+import pandas as pd
+
+out_dir = Path(os.getenv('DATA_MINING_DOC_DIR')) / 'figures'
+
+@click.command('plot:link-elbow')
+def elbow():
+ from sklearn.cluster import KMeans
+
+ filename = 'link_cluster_elbow.png'
+
+ DB = connect()
+ df = DB.query("""
+ SELECT
+ *
+ FROM link_edges
+ """).df()
+ pivot = df.pivot(index='parent_id', columns='child_id', values='links').fillna(0)
+
+ to_plot = []
+ for k in range(2, 15):
+ kmeans = KMeans(n_clusters=k, n_init="auto")
+ kmeans.fit(pivot)
+ label = kmeans.labels_
+ coeff = silhouette_score(pivot, label, metric='euclidean')
+ to_plot.append({'k': k, 'inertia' : kmeans.inertia_, 'coeff': coeff})
+ to_plot = pd.DataFrame(to_plot)
+
+ ax = sns.lineplot(x=to_plot.k, y=to_plot.inertia)
+ ax.set(title="elbow criterion plot of clusters", xlabel="bin size (k)", ylabel="sum of squared distances between centroids/points")
+ plt.savefig(out_dir / filename)
+ plt.close()
+
+ # randomly pick 8
+
+@click.command('plot:link-pca-clusters')
+@click.option('--source', type=click.Choice(['links', 'normalized', 'onehot']), default='links')
+def link_pca_clusters(source):
+
+ filename = f"link_pca_clusters_{source}.png"
+
+ DB = connect()
+ df = DB.query(f"""
+ SELECT
+ c.label as cluster
+ ,p.tld
+ --,b.label as bias
+ ,pca.first
+ ,pca.second
+ ,s.cnt as stories
+ FROM top.publisher_clusters_{source} c
+ JOIN top.publishers p
+ ON c.publisher_id = p.id
+ JOIN
+ (
+ select
+ s.publisher_id
+ ,count(1) as cnt
+ FROM top.stories s
+ GROUP BY
+ s.publisher_id
+ ) s
+ ON s.publisher_id = p.id
+ JOIN top.publisher_pca_{source} pca
+ ON pca.publisher_id = p.id
+ """).df()
+ DB.close()
+
+ ax = sns.scatterplot(x=df['first'], y=df['second'], hue=df['cluster'])
+ ax.set(title=f"pca components vs. clusters ({source})", xlabel="first pca component", ylabel="second pca component")
+ plt.savefig(out_dir / filename)
+
+ # .df().groupby(['cluster', 'bias']).describe()
+
+
+
+
+def test():
+ data_dir = Path(os.getenv('DATA_MINING_DATA_DIR'))
+
+ DB.query("""
+ SELECT
+ p.id as publisher_id
+ ,p.name
+ ,p.tld
+ ,cast(b.bias_id as int) as bias_id
+ ,count(1) as stories
+ FROM publishers p
+ JOIN stories s
+ ON s.publisher_id = p.id
+ JOIN publisher_clusters c
+ ON c.publisher_id = p.id
+ LEFT JOIN publisher_bias b
+ ON b.publisher_id = p.id
+ where bias_id is null
+ group by
+ p.id
+ ,p.name
+ ,p.tld
+ ,b.bias_id
+ ORDER BY count(1) desc
+ """)
+
+ # .df().to_csv(data_dir / 'cluster_publishers.csv', sep="|", index=False)
+ DB.close()
+
diff --git a/src/selection.py b/src/selection.py
new file mode 100644
index 0000000..9c34543
--- /dev/null
+++ b/src/selection.py
@@ -0,0 +1,48 @@
+from data.main import connect
+import pandas as pd
+import numpy as np
+
+DB = connect()
+edges = DB.query("""
+ select
+ *
+ from link_edges
+""").df()
+DB.close()
+
+edges
+
+adj = edges.pivot(index='parent_id', columns='child_id', values='links').fillna(0)
+select_publishers = pd.DataFrame(adj.index.tolist(), columns=['publisher_id'])
+
+DB = connect()
+DB.query("create schema top")
+
+DB.query("""
+ CREATE OR REPLACE TABLE top.publishers AS
+ SELECT
+ p.*
+ FROM publishers p
+ JOIN select_publishers s
+ ON s.publisher_id = p.id
+""")
+
+DB.query("""
+ CREATE OR REPLACE TABLE top.stories AS
+ SELECT
+ s.*
+ FROM stories s
+ JOIN top.publishers p
+ ON s.publisher_id = p.id
+ WHERE year(s.published_at) >= 2006
+ AND year(s.published_at) < 2023
+""")
+
+DB.query("""
+ CREATE OR REPLACE TABLE top.related_stories AS
+ SELECT
+ r.*
+ FROM top.stories s
+ JOIN related_stories r
+ ON s.id = r.parent_id
+""")
diff --git a/src/sentence.py b/src/sentence.py
new file mode 100644
index 0000000..4bf5014
--- /dev/null
+++ b/src/sentence.py
@@ -0,0 +1,138 @@
+from transformers import AutoTokenizer, AutoModel
+import torch
+import torch.nn.functional as F
+from data.main import connect, data_dir
+import os
+from pathlib import Path
+import numpy as np
+import pandas as pd
+from tqdm import tqdm
+import click
+
+#Mean Pooling - Take attention mask into account for correct averaging
+def mean_pooling(model_output, attention_mask):
+ token_embeddings = model_output[0] #First element of model_output contains all token embeddings
+ input_mask_expanded = attention_mask.unsqueeze(-1).expand(token_embeddings.size()).float()
+ return torch.sum(token_embeddings * input_mask_expanded, 1) / torch.clamp(input_mask_expanded.sum(1), min=1e-9)
+
+@click.option('-c', '--chunks', type=int, default=500, show_default=True)
+@click.command("sentence:embed")
+def embed(chunks):
+
+ # Load model from HuggingFace Hub
+ tokenizer = AutoTokenizer.from_pretrained('sentence-transformers/all-MiniLM-L6-v2')
+ model = AutoModel.from_pretrained('sentence-transformers/all-MiniLM-L6-v2')
+
+ # load data
+ DB = connect()
+ table = DB.sql("""
+ select
+ id
+ ,title
+ from stories
+ order by id desc
+ """).df()
+ DB.close()
+
+ # normalize text
+ table['title'] = table['title'].str.normalize('NFKD').str.encode('ascii', errors='ignore').str.decode('utf-8')
+
+
+ chunked = np.array_split(table, chunks)
+ # generate embeddings from list of titles
+ iterator = tqdm(chunked, 'embedding')
+ embeddings = []
+ embedding_ids = []
+ for _, chunk in enumerate(iterator):
+ sentences = chunk['title'].tolist()
+ ids = chunk['id'].tolist()
+ # Tokenize sentences
+ encoded_input = tokenizer(sentences, padding=True, truncation=True, return_tensors='pt')
+ # Compute token embeddings
+ with torch.no_grad():
+ model_output = model(**encoded_input)
+ # Perform pooling
+ output = mean_pooling(model_output, encoded_input['attention_mask'])
+ # Normalize embeddings
+ output = F.normalize(output, p=2, dim=1)
+ embeddings.append(output)
+ embedding_ids.append(ids)
+
+ embeddings = np.concatenate(embeddings)
+ ids = np.concatenate(embedding_ids)
+
+ # save embeddings
+ save_to = data_dir() / 'embeddings.npy'
+ np.save(save_to, embeddings)
+ print(f"embeddings saved: {save_to}")
+
+ # save ids
+ save_to = data_dir() / 'embedding_ids.npy'
+ np.save(save_to, ids)
+ print(f"ids saved: {save_to}")
+
+
+@click.command('sentence:create-pca-table')
+def create_table():
+ from sklearn import linear_model
+ data_path = Path(os.getenv('DATA_MINING_DATA_DIR'))
+
+ embeddings = np.load(data_path / 'embeddings.npy')
+ embedding_ids = np.load(data_path / 'embedding_ids.npy')
+ ids = pd.DataFrame(embedding_ids, columns=['story_id']).reset_index()
+ DB = connect()
+
+ data = DB.query("""
+ SELECT
+ ids.index
+ ,s.id
+ ,b.ordinal
+ FROM ids
+ JOIN top.stories s
+ ON ids.story_id = s.id
+ JOIN top.publisher_bias pb
+ ON pb.publisher_id = s.publisher_id
+ JOIN bias_ratings b
+ ON b.id = pb.bias_id
+ """).df()
+
+ x = embeddings[data['index']]
+ y = data['ordinal'].to_numpy().reshape(-1, 1)
+
+ reg = linear_model.LinearRegression()
+
+ reg.fit(x, y)
+
+ reg.coef_.shape
+
+@click.command('sentence:create-svm-table')
+def create_svm_table():
+ from sklearn import svm
+ data_path = Path(os.getenv('DATA_MINING_DATA_DIR'))
+
+ embeddings = np.load(data_path / 'embeddings.npy')
+ embedding_ids = np.load(data_path / 'embedding_ids.npy')
+ ids = pd.DataFrame(embedding_ids, columns=['story_id']).reset_index()
+
+ DB = connect()
+ data = DB.query("""
+ SELECT
+ ids.index
+ ,s.id
+ ,b.ordinal
+ FROM ids
+ JOIN top.stories s
+ ON ids.story_id = s.id
+ JOIN top.publisher_bias pb
+ ON pb.publisher_id = s.publisher_id
+ JOIN bias_ratings b
+ ON b.id = pb.bias_id
+ """).df()
+
+ x = embeddings[data['index']]
+ #y = data['ordinal'].to_numpy().reshape(-1, 1)
+ y = data['ordinal']
+
+ clf = svm.SVC()
+ pred = clf.fit(x, y)
+
diff --git a/src/train/__init__.py b/src/train/__init__.py
new file mode 100644
index 0000000..4519cec
--- /dev/null
+++ b/src/train/__init__.py
@@ -0,0 +1,5 @@
+import train.main
+
+__all__ = [
+ 'main'
+]
diff --git a/src/train/dataset.py b/src/train/dataset.py
new file mode 100644
index 0000000..15bfd47
--- /dev/null
+++ b/src/train/dataset.py
@@ -0,0 +1,38 @@
+from torch.utils.data import Dataset
+from data.main import connect, data_dir
+from bias import label_to_int
+import numpy as np
+import pandas as pd
+
+class NewsDataset(Dataset):
+ def __init__(self):
+ self.embeddings = np.load(data_dir() / 'embeddings.npy')
+ embedding_ids = pd.DataFrame(np.load(data_dir() / 'embedding_ids.npy'), columns=['id']).reset_index()
+
+ DB = connect()
+ query = """
+ SELECT
+ s.id
+ ,b.label
+ ,count(1) over (partition by publisher_id) as stories
+ FROM stories s
+ JOIN publisher_bias b
+ ON b.id = s.publisher_id
+ WHERE b.label != 'allsides'
+ """
+ data = DB.sql(query).df()
+ DB.close()
+
+ data['label'] = data['label'].apply(lambda x: label_to_int(x))
+ data = data.merge(embedding_ids)
+ self.data = data
+
+ def __len__(self):
+ return len(self.data)
+
+ def __getitem__(self, idx):
+ row = self.data.iloc[idx]
+ y = row['label']
+ # x = np.concatenate((self.embeddings[row['index']], [row['stories']])).astype(np.float32)
+ x = self.embeddings[row['index']]
+ return x, y
diff --git a/src/train/main.py b/src/train/main.py
new file mode 100644
index 0000000..dbad152
--- /dev/null
+++ b/src/train/main.py
@@ -0,0 +1,132 @@
+import click
+from tqdm import tqdm
+from enum import Enum, auto
+from dotenv import load_dotenv
+import os
+
+import torch
+from torch import nn
+from torch import optim
+from torch.utils.data import DataLoader
+from accelerate import Accelerator
+
+from train.dataset import NewsDataset
+from train.model import Classifier
+#from model.linear import LinearClassifier
+
+class Stage(Enum):
+ TRAIN = auto()
+ DEV = auto()
+
+@click.command('train:main')
+def main():
+ dev_after = 20
+ visible_devices = None
+ lr = 1e-4
+ epochs = 10
+ debug = False
+ torch.manual_seed(0)
+ num_workers = 0
+
+ embedding_length = int(os.getenv('EMBEDDING_LENGTH', 384))
+
+ dataset = NewsDataset()
+ trainset, devset = torch.utils.data.random_split(dataset, [0.8, 0.2])
+ batch_size = 512
+ trainloader = DataLoader(trainset, batch_size=batch_size, shuffle=True, num_workers=num_workers, drop_last=True)
+ devloader = DataLoader(devset, shuffle=False, num_workers=num_workers)
+ accelerator = Accelerator()
+ model = Classifier(embedding_length=embedding_length, classes=5)
+
+ # it's possible to control which GPUs the process can see using an environmental variable
+ if visible_devices:
+ os.environ['CUDA_VISIBLE_DEVICES'] = visible_devices
+ if debug:
+ os.environ['CUDA_LAUNCH_BLOCKING'] = '1'
+ #accelerator.log({"message" :"debug enabled"})
+
+ criterion = torch.nn.CrossEntropyLoss()
+ optimizer = torch.optim.Adam(model.parameters(), lr=lr)
+
+ # wrap objects with accelerate
+ model, optimizer, trainloader, devloader = accelerator.prepare(model, optimizer, trainloader, devloader)
+ def run():
+ """runner for training and valdating"""
+ running_loss = 0.0
+ # set the model to train model
+ model.train() if stage == Stage.TRAIN else model.eval()
+ dataloader = trainloader if stage == Stage.TRAIN else devloader
+ desc = 'train epoch' if stage == Stage.TRAIN else 'dev epoch'
+ if debug:
+ ...
+
+ # Make sure there are no leftover gradients before starting training an epoch
+ optimizer.zero_grad()
+
+ for batch, (x, y) in enumerate(tqdm(dataloader, desc=desc)):
+ pred_y = model(x) # Forward pass through model
+ loss = criterion(pred_y, y)
+ running_loss += loss # Increment running loss
+ # Only update model weights on training
+ if stage == Stage.TRAIN:
+ accelerator.backward(loss) # Increment gradients within model by sending loss backwards
+ optimizer.step() # Update model weights
+ optimizer.zero_grad() # Reset gradients to 0
+ return running_loss / len(dataloader)
+
+
+ for epoch in range(epochs):
+ if (epoch - 1) % dev_after == 0:
+ stage = Stage.DEV
+ log = run()
+ print(f"dev loss: {log}")
+ else:
+ stage = Stage.TRAIN
+ log = run()
+ print(f"train loss: {log}")
+
+ breakpoint()
+ from data.main import data_dir, connect
+ import numpy as np
+ import pandas as pd
+ from bias import int_to_label
+
+ embeddings = dataset.embeddings
+ embedding_ids = dataset.data
+
+ DB = connect()
+ query = """
+ SELECT
+ s.id
+ ,title
+ ,p.name
+ ,count(1) over (partition by publisher_id) as stories
+ FROM stories s
+ JOIN publishers p
+ on p.id = s.publisher_id
+ WHERE s.publisher_id NOT IN (
+ SELECT
+ id
+ FROM publisher_bias b
+ )
+ """
+ data = DB.sql(query).df()
+ embeddings = np.load(data_dir() / 'embeddings.npy')
+ embedding_ids = pd.DataFrame(np.load(data_dir() / 'embedding_ids.npy'), columns=['id']).reset_index()
+
+
+ for i in range(10):
+ embedding = embeddings[embedding_ids[embedding_ids['id'] == data.iloc[i]['id']]['index']]
+ title = data.iloc[i]['title']
+ publisher = data.iloc[i]['name']
+ class_pred = nn.functional.softmax( model(torch.tensor(embedding))).detach()
+ class_id = int(torch.argmax(nn.functional.softmax( model(torch.tensor(embedding))).detach()))
+ print(f"{publisher}: {int_to_label(class_id)} - \"{title}\"")
+
+ embedding_ids['id'] == data.iloc[0]['id']
+ embedding_ids[embedding_ids['id'] == data.iloc[0]['id']]
+ embedding = embeddings[embedding_ids[embedding_ids['id'] == data.iloc[0]['id']]['index']]
+ title
+ publisher
+
+ model().get_last_layer(torch.tensor(embedding))
diff --git a/src/train/model.py b/src/train/model.py
new file mode 100644
index 0000000..b9faed4
--- /dev/null
+++ b/src/train/model.py
@@ -0,0 +1,28 @@
+from torch import nn
+
+class Classifier(nn.Module):
+ def __init__(self, embedding_length: int, classes: int):
+ super().__init__()
+ out_len = 16
+ self.stack = nn.Sequential(
+ nn.Linear(embedding_length, 256),
+ nn.ReLU(),
+ nn.Linear(256, 256),
+ nn.ReLU(),
+ nn.Linear(256, 64),
+ nn.ReLU(),
+ nn.Linear(64, 64),
+ nn.ReLU(),
+ nn.Linear(64, out_len),
+ nn.ReLU(),
+ )
+ self.logits = nn.Linear(out_len, classes)
+
+ def forward(self, x):
+ x = self.stack(x)
+ self.last_hidden_layer = x.detach()
+ return self.logits(x)
+
+ def get_last_layer(self, x):
+ x = self.stack(x)
+ return x
diff --git a/src/word.py b/src/word.py
index dc408d5..36f5244 100644
--- a/src/word.py
+++ b/src/word.py
@@ -1,7 +1,7 @@
import click
from transformers import AutoTokenizer, RobertaModel
import numpy as np
-from data import Data, from_db, connect, data_dir
+from data.main import Data, from_db, connect, data_dir
from tqdm import tqdm
import torch
from pathlib import Path